Microscopy analysis, reimagined for the cloud
Open-source platform that brings powerful segmentation, machine learning, and quantitative analysis to your microscopy data — all from your browser.
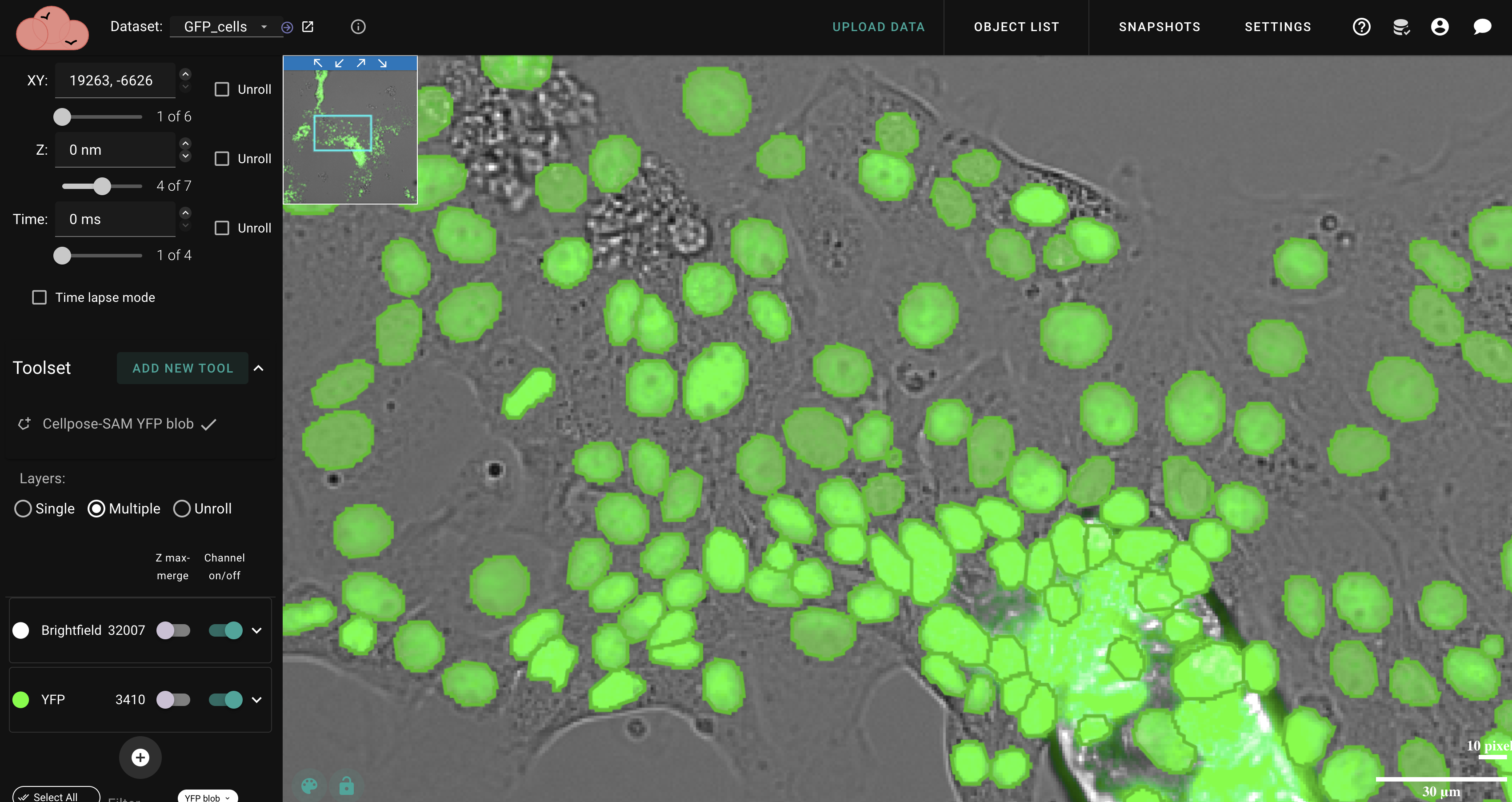
Open-source platform that brings powerful segmentation, machine learning, and quantitative analysis to your microscopy data — all from your browser.

Access your work from anywhere. Analyze vast datasets with advanced machine learning — no installation, no local compute limits, just your browser.
Put your hands on your data in completely new ways. Intuitive tools make complex image analysis accessible and — dare we say — enjoyable.
Our code is freely available for scrutiny, modification, and improvement. Run your own server, contribute to the project, or build on top of it.
Segment cells, nuclei, and structures with Cellpose, Piscis, and SAM. Fine-tune results with intuitive manual tools.
Detect and quantify fluorescent spots with high accuracy — perfect for RNA FISH, protein aggregates, and spatial analysis.
Track cells and objects across time-lapse images. Maintain identity through divisions for lineage tracing and cell cycle analysis.
Measure morphology, fluorescence intensity, and spatial relationships. Analyze distributions across experimental conditions.
Process 3D stacks and multiple fluorescent channels. Investigate colocalization, proximity, and interactions between components.
Process large datasets with automated workflows. Integrate machine learning models or train new ones on your own data.
NimbusImage was developed over 15 years in the Raj Lab at the University of Pennsylvania in collaboration with Kitware, leaders in open-source scientific software. Read about the platform in Nature Methods.
Whether you're a solo researcher or running an imaging core, there's a plan that fits.
NimbusImage runs in any modern browser, but we also provide a standalone desktop app that isolates your workspace and delivers better performance for intensive sessions.
Isolate NimbusImage from browser tabs for smoother rendering on large datasets.
Native builds for Windows, macOS (Apple Silicon), and Linux.
Same cloud account, same data. Switch between browser and app seamlessly.
NimbusImage is built on open-source principles. Our core technology is developed transparently, and the commercial deployment through CytoPixel Software provides a stable, supported experience.
We are deeply committed to creating accessible, robust tools that help researchers unlock the full potential of their imaging data. Our primary goal is to create tools that assist and delight.
University of Pennsylvania. 15+ years of molecular imaging research powering NimbusImage.
Industry leaders in open-source software. Developers of Girder and ITK.
Commercial deployment and support for the NimbusImage platform.
Sign up and start analyzing in minutes. No installation, no credit card.